# Load CSV file from github
H32Metadata <- read_csv("https://raw.githubusercontent.com/cincibrainlab/vhtp/main/chanfiles/NeuroNexusMouseEEGv2H32_Map.csv", col_types = cols())
# View the first 10 rows of the metadata
tb1 <- H32Metadata %>%
head(5) %>%
kable()Overview
For alignment, here are the exact description of the equipment we are using from our purchase order:
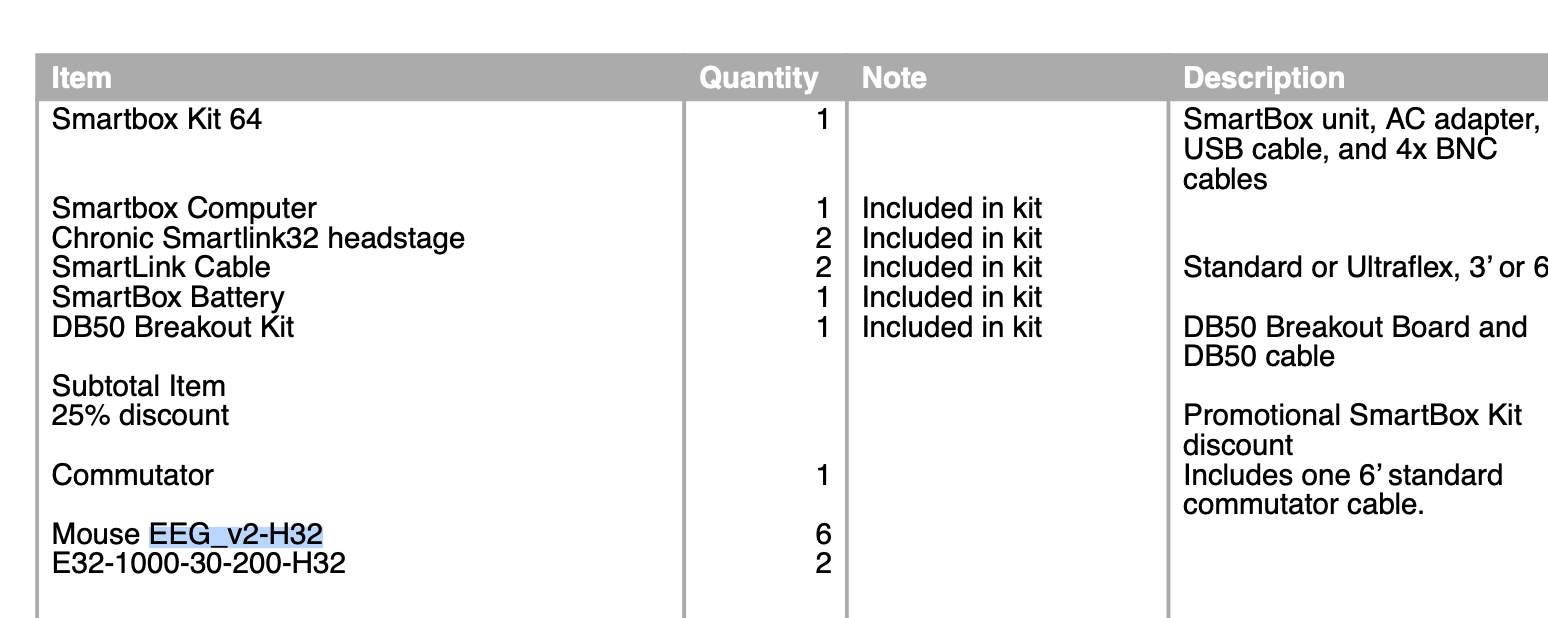

This protocol describes how to map the location of the Mouse EEG v2 H32 Probe with the OM Amplifier Adapter. For better or worse, like all custom probes mapping must be confirmed manually.
Probe Layout
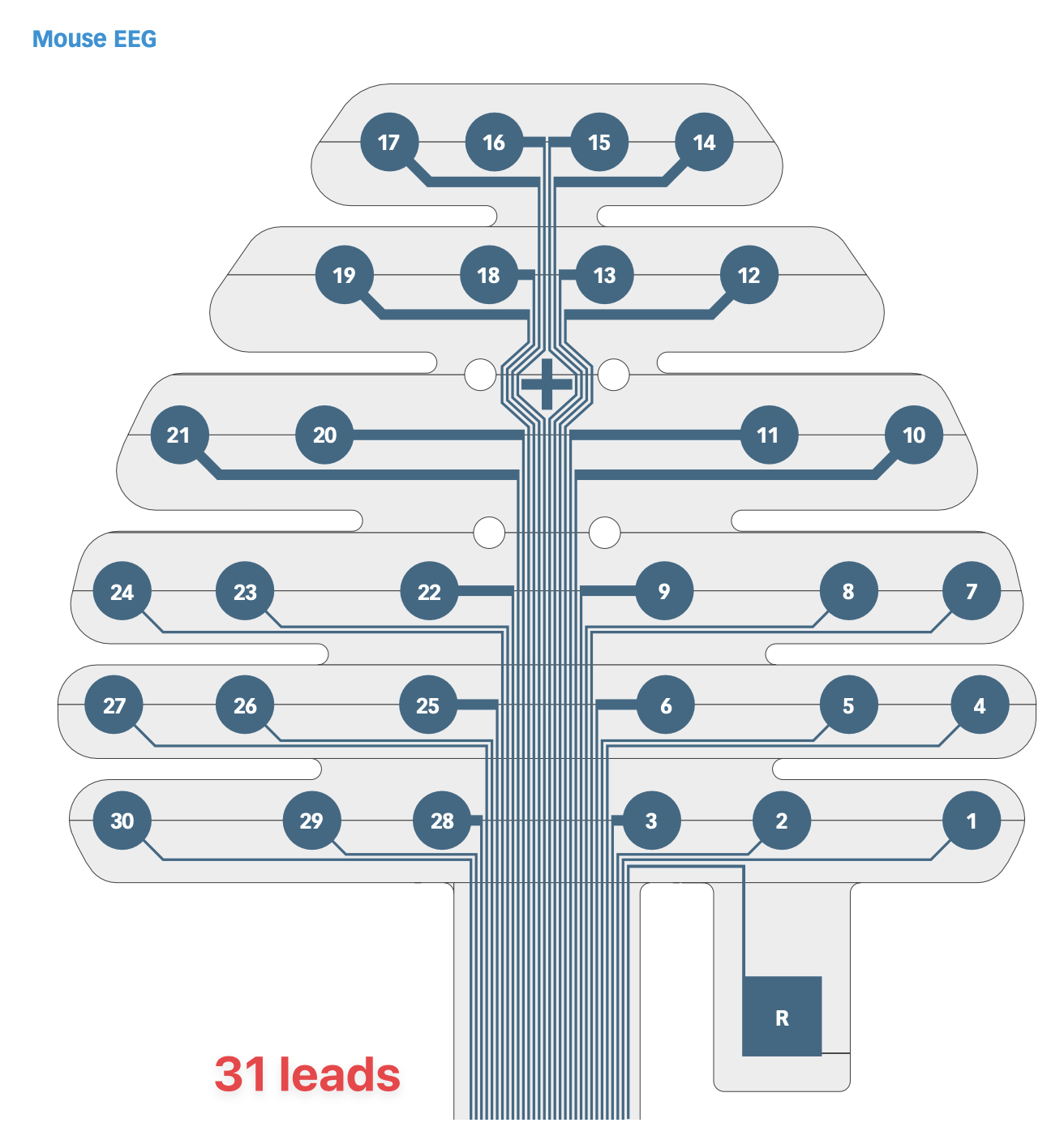

This image is from the “bottom up” so the reference electrode is actually on the bottom left following implantation:
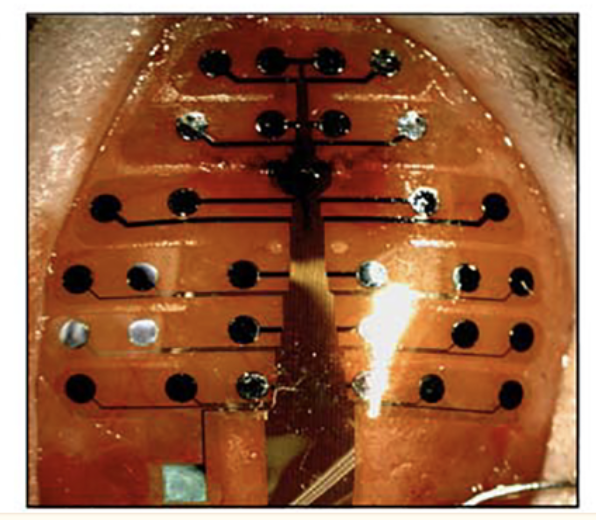
Per Neuronexus the recordings are stored in the following format:
Radiens saves primary time-series data sets (e.g., neural recordings) in a simple, open format. Each dataset (i.e. recording) is saved in a file set named ‘XDAT’ that consists of three files: (i) flat binary file containing the matrix of all sample values (typical in uV) across the multiple channels, (ii) flat binary file containing all timestamps, (iii) json text file containing enriched metadata. link
View the raw signal data using MATLAB
A native MATLAB function is available to import the XDAT dataset. The function is available https://github.com/nzgurel/AllegoProcessing_Matlab/tree/main with an example import script here.
The raw data array can be plotted using the EEGLAB functio eegplot:
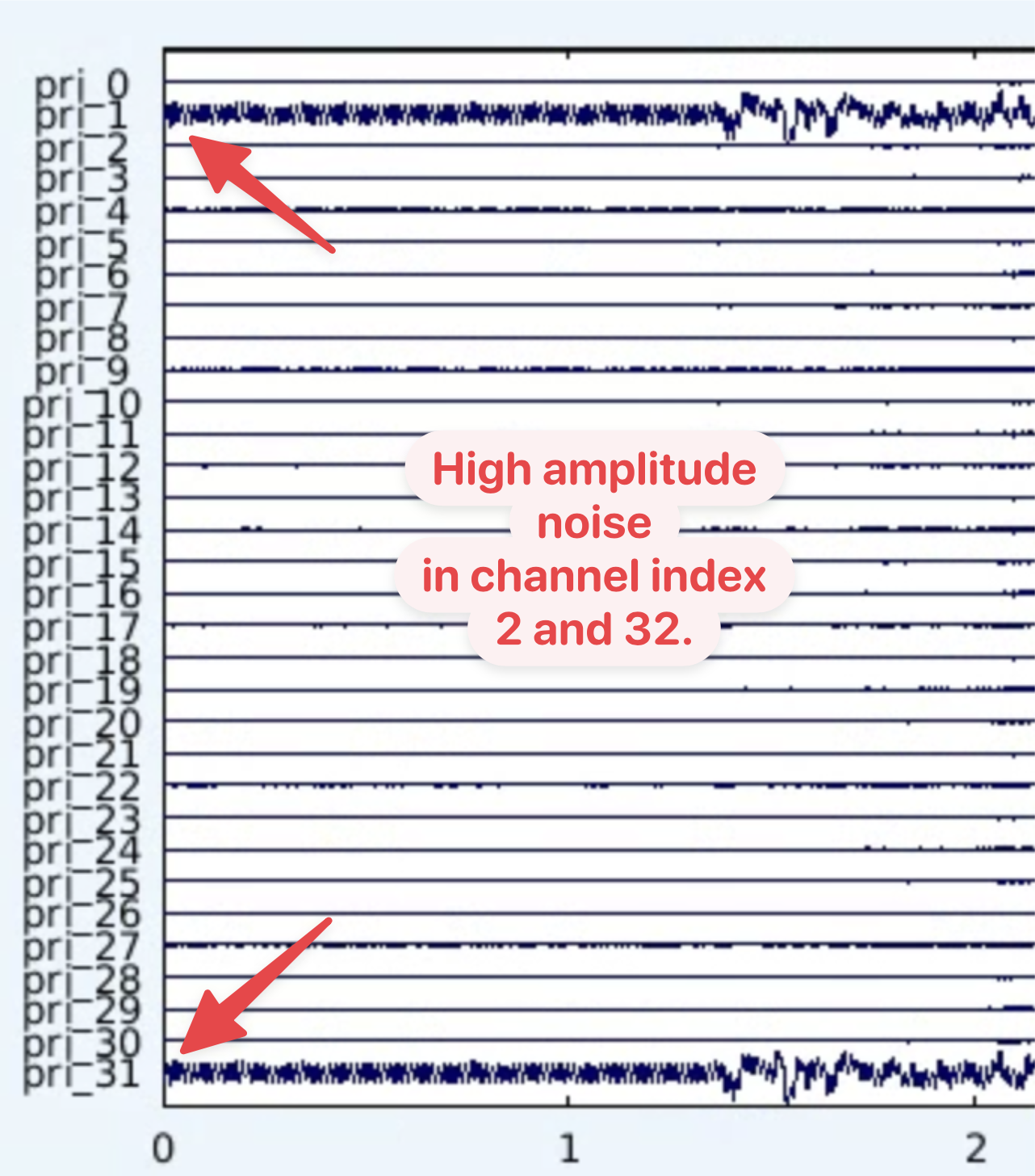
Here we can see that channels pri_1 and pri_31 are not connected to recording sites. As the naming index starts with 0, following import into MATLAB these will be channels indexes 1 and 32.
View and export the metata data using MATLAB
We can import metadata from the json file which contains infomation about the probe layout and recording and export it as a CSV file for further analysis.
% MATLAB code to extract metadata from XDAT dataset
str = fileread([xdat_filename '.xdat.json']);
jsonData = jsondecode(str);
% extract metadata for probe mapping
H32Metadata = jsonData.sapiens_base.biointerface_map;
% remove non-array fields
H32Metadata = rmfield(H32Metadata, 'samp_freq');
H32Metadata = rmfield(H32Metadata, 'sig_units');
% save metadata as CSV
writetable(struct2table(H32Metadata),'NeuroNexusMouseEEGv2H32_Map.csv');For convenience, I have hosted the resulting CSV file on github for futher analysis.
Examining the Probe Metadata
We will evalute the metadata in R to more easily plot and wrangle the data. First, we will load the CSV file from github and view the first 5 rows of the metadata.
Table 1: Table of XDAT dataset metadata for the Mouse EEG v2 H32 Probe.
| chan_name | chan_type | ntv_chan_idx | ntv_chan_name | sys_chan_idx | site_shape | port | absolute_idx | scs_to_tcs_y | site_lim_x_min | site_lim_z_max | site_ctr_y | headstage_id | site_ctr_z | site_ctr_x | site_ctr_tcs_x | site_lim_x_max | probe_id | site_ctr_tcs_y | site_num | sensor_units | scs_to_tcs_z | is_audio | is_audio_left | is_audio_right | is_selected | site_lim_y_min | site_lim_y_max | scs_to_tcs_x | site_lim_z_min | dsrc_uid | sensor_uid | site_ctr_tcs_z | color_grp_idx |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| pri_0 | ai0 | 0 | port_A_0 | 0 | circle | A | 4 | 0 | 674.114 | 0 | 404.99961 | hstg__passthrough_32 | 0 | 699.1136 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 1 | uV | 0 | 0 | 0 | 0 | 1 | 430 | 380 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_1 | ai0 | 1 | port_A_1 | 1 | circle | A | 5 | 0 | 674.114 | 0 | 223.99961 | hstg__passthrough_32 | 0 | 699.1136 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 2 | uV | 0 | 0 | 0 | 0 | 1 | 249 | 199 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_2 | ai0 | 2 | port_A_2 | 2 | circle | A | 12 | 0 | 674.114 | 0 | 99.99961 | hstg__passthrough_32 | 0 | 699.1136 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 3 | uV | 0 | 0 | 0 | 0 | 1 | 125 | 75 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 3 |
| pri_3 | ai0 | 3 | port_A_3 | 3 | circle | A | 6 | 0 | 564.114 | 0 | 412.99961 | hstg__passthrough_32 | 0 | 589.1136 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 4 | uV | 0 | 0 | 0 | 0 | 1 | 438 | 388 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_4 | ai0 | 4 | port_A_4 | 4 | circle | A | 7 | 0 | 564.114 | 0 | 287.99961 | hstg__passthrough_32 | 0 | 589.1136 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 5 | uV | 0 | 0 | 0 | 0 | 1 | 313 | 263 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
Evaluating the Probe Metadata
The metadata hasa several curosities that suggest using caution before incorporating it into your workflow.
- The Murine EEG v2 H32 Probe has 30 channels, not 32 (pri_0 to pri_31), as indicated by the metadata.
- 2D coordinate data is provided for the first 30 channels (pri_0 to pri_29), but not the last two (pri_30 and pri_31).
- There is no indication that channels pri_1 and pri_31 are not connected to recording sites.
This can be seen more clearly by plotting out the channels and their coordinates on a 2d scatter plot. Indeed, this layout matches the mouse 30 lead layout above.
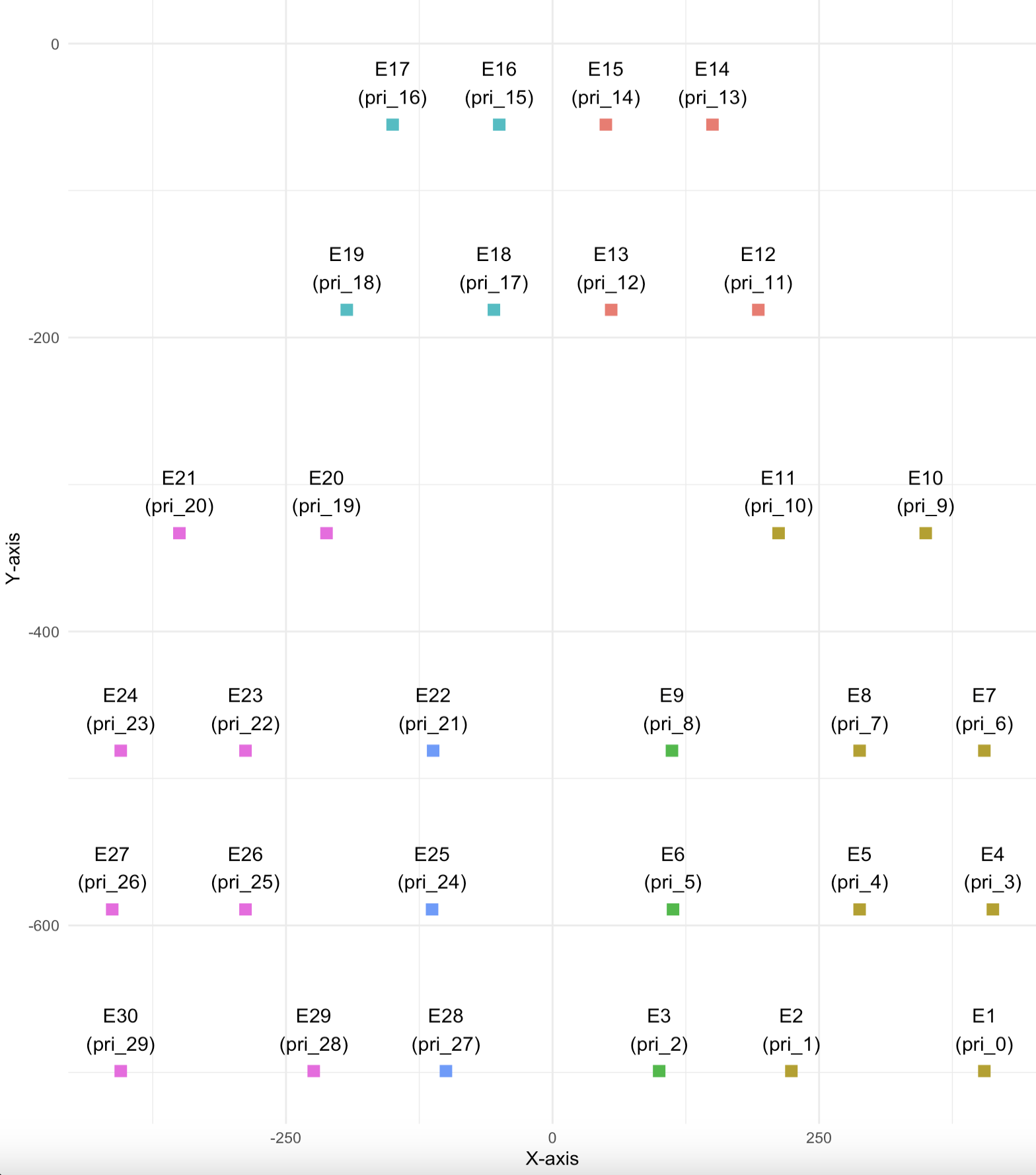
This suggests that the metadata has the correct layout information but has not been adjusted foe the two unconnected channels. I don’t understand why this is the case, but I wil be reaching out to Neuronexus to confirm and will post their reponse.
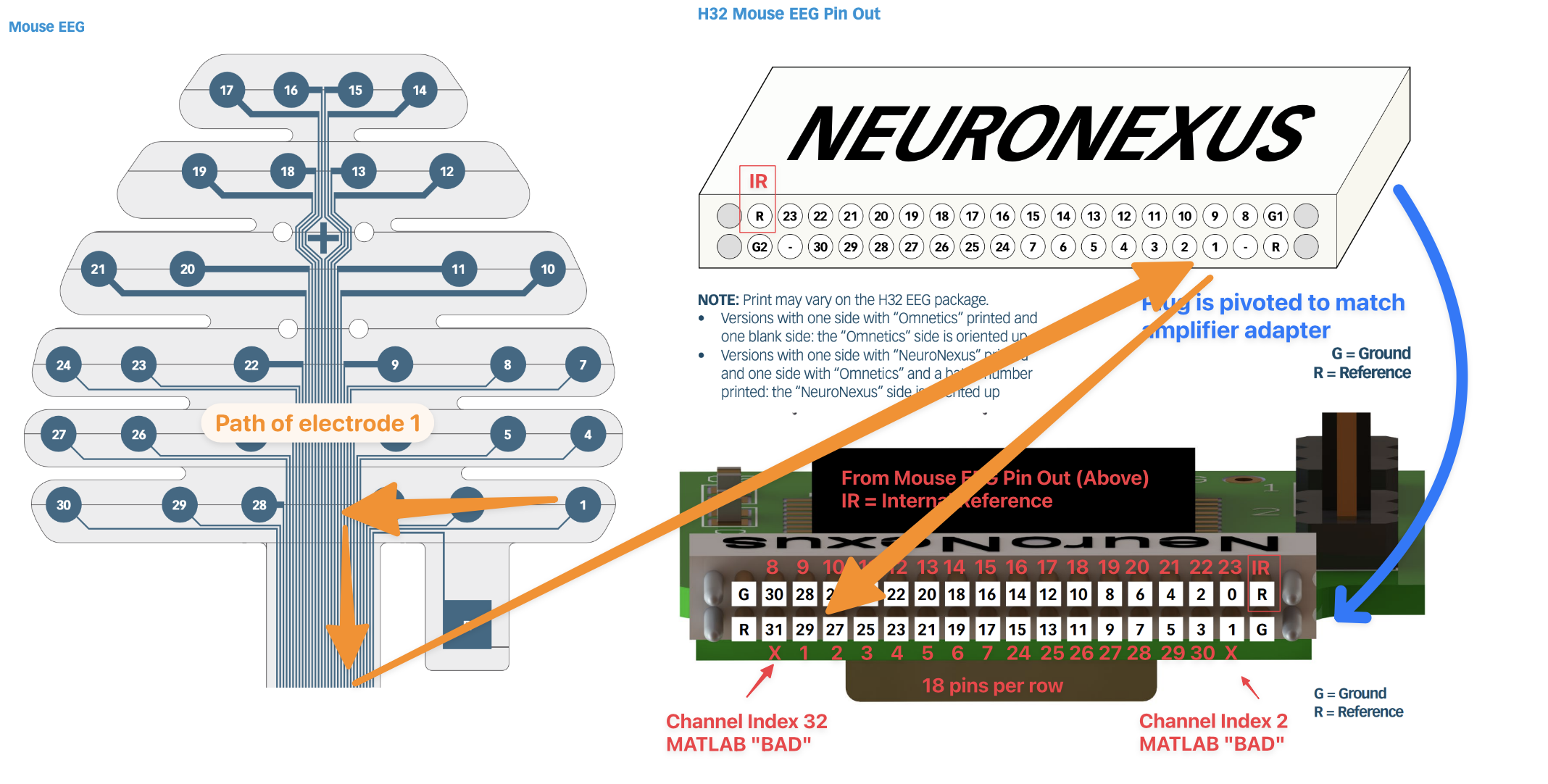
Tracing the Probe electrodes to the amplifier adapter
The probe is connected to the headstage using the Ometics Amplifier Adapter (NPD-36-AA-GS). This connector has 36 ports and 4 guide posts.
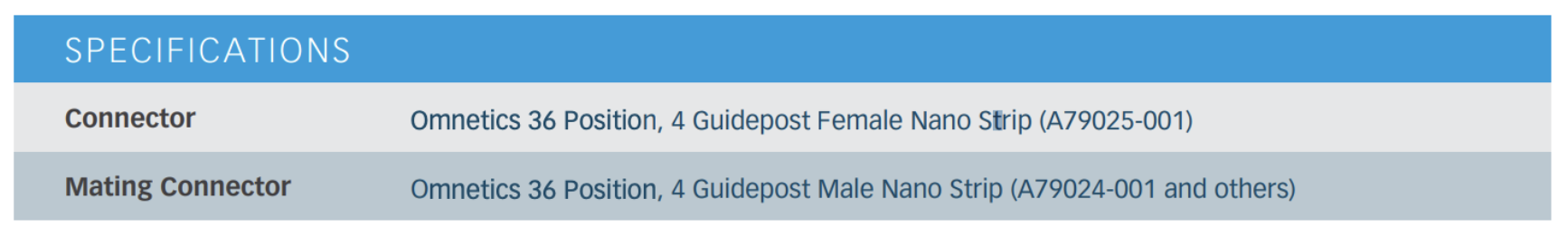

These layouts are not necessarily easy to find in one place so I have included them stiched together here:
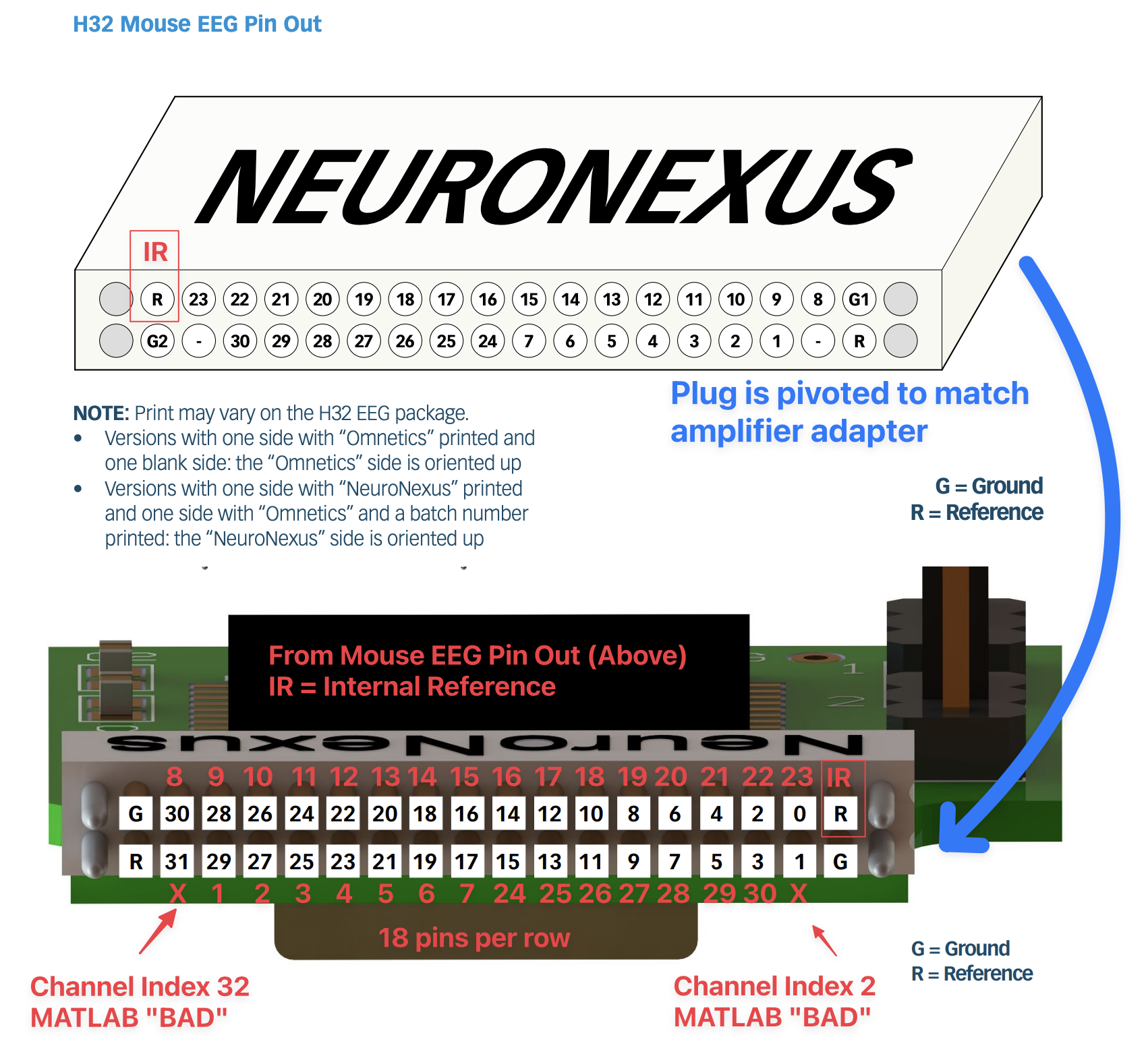
The Neuronexus EEG Pin Out must be rotated to match the Amplifier Adapter Pin Out. To make it easier to visualize the matching channels, I have added red EEG channel number labels to the Amplifier Adapter Pin Out. Importantly, you can see that the Mouse 30-lead EEG leaves two channels unconnected (pri_1 and pri_31). This now clearly visible in the adapter image.
Per the documentation the internal reference electrode is connected to R1. This corresponds to the “R” pin adjacent to the “0” pin once you account for the rotation of the pin out.
Mapping the Probe to the Amplifier Adapter Stage 1
The first step is to create a simple lookup table that disregards all metadata but the amplifier pin out (pri_) and the EEG probe channels (based on the connection).
This mapping will correctly pair each EEG channel to the correct signal entering the amplifier, but will not have the correct 30 lead metadata till after we remove the unconnected channels.
mea_to_hs_map <- c(
"E1" = 29,
"E2" = 27,
"E3" = 25,
"E4" = 23,
"E5" = 21,
"E6" = 19,
"E7" = 17,
"E8" = 30,
"E9" = 28,
"E10" = 26,
"E11" = 24,
"E12" = 22,
"E13" = 20,
"E14" = 18,
"E15" = 16,
"E16" = 14,
"E17" = 12,
"E18" = 10,
"E19" = 8,
"E20" = 6,
"E21" = 4,
"E22" = 2,
"E23" = 0,
"E24" = 15,
"E25" = 13,
"E26" = 11,
"E27" = 9,
"E28" = 7,
"E29" = 5,
"E30" = 3
)
# Create a dataframe from the named list
# convert this to a dataframe
mea_to_hs_map_df <- data.frame(mea_to_hs_map) %>%
rownames_to_column(var = "mea") %>%
rename(hs_idx = mea_to_hs_map) %>%
as_tibble() %>%
mutate(headset = paste0("pri_", hs_idx)) %>%
rename(mea_formatted_name = mea,
og_amplifier_pin = headset,
amplifier_index = hs_idx)
mea_to_hs_map_df %>% head(3) %>% kable()| mea_formatted_name | amplifier_index | og_amplifier_pin |
|---|---|---|
| E1 | 29 | pri_29 |
| E2 | 27 | pri_27 |
| E3 | 25 | pri_25 |
write_csv(mea_to_hs_map_df, "MouseEEGv2H32_Import_Stage1.csv")The final table has the final formatted MEA channel name (E1 to E30) and the headset channel name (pri_0 to pri_29). The original index (starting with 1) is also included. We will save this file to use in MATLAB during import.
The file can be downloaded here.
We can now accurately map which MEA channels correspond to which amplifier channels. The bad channels (pri_1 and pri_31) now make more sense as you can follow the mapping.
Where is the reference channel?

As of now, we cannot find data for the reference channel (R1) either in the metadata or from the raw read of the signal data. We will need to reach out to Neuronexus to confirm location of reference channel. Since the reference channel can either by an external wire or an internal channel determined by a physical jumper wire, we would not expect the metadata to contain this information inherently. However, the actual signal data should be present.
Mapping the Probe to the Amplifier Adapter Stage 2
Now that the mapping is complete, we can remove the unconnected channels and add the metadata to the mapping.
As you can see from the manual plot above, pri_0 contains the correct metadata for electrode E1. However, in actuality E1 data is routed through pri_29.
Thus, the metadata is now more accurately used as a lookup table for the probe channel and uncoupled from the amplifier channel. If we assigned the metadata for pri_29 to E1, the location would be incorrect. Let’s create a stage2 mapping that has the correct metadata (primarily coordinates) for each electrode.
# read metadata csv from github
mea_metadata <- "https://raw.githubusercontent.com/cincibrainlab/vhtp/main/chanfiles/NeuroNexusMouseEEGv2H32_Map.csv"
stage2 <- read_csv(mea_metadata, col_types = cols()) %>%
filter(sys_chan_idx < 30) %>%
mutate(mea_formatted_name = paste0("E", sys_chan_idx+1)) %>%
left_join(mea_to_hs_map_df %>% select(mea_formatted_name,og_amplifier_pin), by = "mea_formatted_name") %>%
relocate(mea_formatted_name, .after = chan_name) %>%
relocate(og_amplifier_pin, .after = mea_formatted_name)
stage2 %>% kable()| chan_name | mea_formatted_name | og_amplifier_pin | chan_type | ntv_chan_idx | ntv_chan_name | sys_chan_idx | site_shape | port | absolute_idx | scs_to_tcs_y | site_lim_x_min | site_lim_z_max | site_ctr_y | headstage_id | site_ctr_z | site_ctr_x | site_ctr_tcs_x | site_lim_x_max | probe_id | site_ctr_tcs_y | site_num | sensor_units | scs_to_tcs_z | is_audio | is_audio_left | is_audio_right | is_selected | site_lim_y_min | site_lim_y_max | scs_to_tcs_x | site_lim_z_min | dsrc_uid | sensor_uid | site_ctr_tcs_z | color_grp_idx |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| pri_0 | E1 | pri_29 | ai0 | 0 | port_A_0 | 0 | circle | A | 4 | 0 | 674.114 | 0 | 404.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 1 | uV | 0 | 0 | 0 | 0 | 1 | 430 | 380 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_1 | E2 | pri_27 | ai0 | 1 | port_A_1 | 1 | circle | A | 5 | 0 | 674.114 | 0 | 223.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 2 | uV | 0 | 0 | 0 | 0 | 1 | 249 | 199 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_2 | E3 | pri_25 | ai0 | 2 | port_A_2 | 2 | circle | A | 12 | 0 | 674.114 | 0 | 99.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 3 | uV | 0 | 0 | 0 | 0 | 1 | 125 | 75 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 3 |
| pri_3 | E4 | pri_23 | ai0 | 3 | port_A_3 | 3 | circle | A | 6 | 0 | 564.114 | 0 | 412.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 4 | uV | 0 | 0 | 0 | 0 | 1 | 438 | 388 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_4 | E5 | pri_21 | ai0 | 4 | port_A_4 | 4 | circle | A | 7 | 0 | 564.114 | 0 | 287.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 5 | uV | 0 | 0 | 0 | 0 | 1 | 313 | 263 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_5 | E6 | pri_19 | ai0 | 5 | port_A_5 | 5 | circle | A | 13 | 0 | 564.114 | 0 | 112.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 6 | uV | 0 | 0 | 0 | 0 | 1 | 138 | 88 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 3 |
| pri_6 | E7 | pri_17 | ai0 | 6 | port_A_6 | 6 | circle | A | 8 | 0 | 456.114 | 0 | 404.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 7 | uV | 0 | 0 | 0 | 0 | 1 | 430 | 380 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_7 | E8 | pri_30 | ai0 | 7 | port_A_7 | 7 | circle | A | 9 | 0 | 456.114 | 0 | 287.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 8 | uV | 0 | 0 | 0 | 0 | 1 | 313 | 263 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_8 | E9 | pri_28 | ai0 | 8 | port_A_8 | 8 | circle | A | 14 | 0 | 456.114 | 0 | 111.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 9 | uV | 0 | 0 | 0 | 0 | 1 | 137 | 87 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 3 |
| pri_9 | E10 | pri_26 | ai0 | 9 | port_A_9 | 9 | circle | A | 10 | 0 | 308.114 | 0 | 349.99961 | hstg__passthrough_32 | 0 | 333.11361 | 0 | 358.114 | probe__eeg_mouse_1x30 | 0 | 10 | uV | 0 | 0 | 0 | 0 | 1 | 375 | 325 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_10 | E11 | pri_24 | ai0 | 10 | port_A_10 | 10 | circle | A | 11 | 0 | 308.114 | 0 | 211.99961 | hstg__passthrough_32 | 0 | 333.11361 | 0 | 358.114 | probe__eeg_mouse_1x30 | 0 | 11 | uV | 0 | 0 | 0 | 0 | 1 | 237 | 187 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 2 |
| pri_11 | E12 | pri_22 | ai0 | 11 | port_A_11 | 11 | circle | A | 0 | 0 | 156.114 | 0 | 192.99961 | hstg__passthrough_32 | 0 | 181.11439 | 0 | 206.114 | probe__eeg_mouse_1x30 | 0 | 12 | uV | 0 | 0 | 0 | 0 | 1 | 218 | 168 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 1 |
| pri_12 | E13 | pri_20 | ai0 | 12 | port_A_12 | 12 | circle | A | 1 | 0 | 156.114 | 0 | 54.99961 | hstg__passthrough_32 | 0 | 181.11439 | 0 | 206.114 | probe__eeg_mouse_1x30 | 0 | 13 | uV | 0 | 0 | 0 | 0 | 1 | 80 | 30 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 1 |
| pri_13 | E14 | pri_18 | ai0 | 13 | port_A_13 | 13 | circle | A | 2 | 0 | 30.114 | 0 | 149.99961 | hstg__passthrough_32 | 0 | 55.11439 | 0 | 80.114 | probe__eeg_mouse_1x30 | 0 | 14 | uV | 0 | 0 | 0 | 0 | 1 | 175 | 125 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 1 |
| pri_14 | E15 | pri_16 | ai0 | 14 | port_A_14 | 14 | circle | A | 3 | 0 | 30.114 | 0 | 49.99961 | hstg__passthrough_32 | 0 | 55.11439 | 0 | 80.114 | probe__eeg_mouse_1x30 | 0 | 15 | uV | 0 | 0 | 0 | 0 | 1 | 75 | 25 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 1 |
| pri_15 | E16 | pri_14 | ai0 | 15 | port_A_15 | 15 | circle | A | 15 | 0 | 30.114 | 0 | -49.99961 | hstg__passthrough_32 | 0 | 55.11439 | 0 | 80.114 | probe__eeg_mouse_1x30 | 0 | 16 | uV | 0 | 0 | 0 | 0 | 1 | -25 | -75 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 4 |
| pri_16 | E17 | pri_12 | ai0 | 16 | port_A_16 | 16 | circle | A | 16 | 0 | 30.114 | 0 | -149.99961 | hstg__passthrough_32 | 0 | 55.11439 | 0 | 80.114 | probe__eeg_mouse_1x30 | 0 | 17 | uV | 0 | 0 | 0 | 0 | 1 | -125 | -175 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 4 |
| pri_17 | E18 | pri_10 | ai0 | 17 | port_A_17 | 17 | circle | A | 17 | 0 | 156.114 | 0 | -54.99961 | hstg__passthrough_32 | 0 | 181.11439 | 0 | 206.114 | probe__eeg_mouse_1x30 | 0 | 18 | uV | 0 | 0 | 0 | 0 | 1 | -30 | -80 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 4 |
| pri_18 | E19 | pri_8 | ai0 | 18 | port_A_18 | 18 | circle | A | 18 | 0 | 156.114 | 0 | -192.99961 | hstg__passthrough_32 | 0 | 181.11439 | 0 | 206.114 | probe__eeg_mouse_1x30 | 0 | 19 | uV | 0 | 0 | 0 | 0 | 1 | -168 | -218 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 4 |
| pri_19 | E20 | pri_6 | ai0 | 19 | port_A_19 | 19 | circle | A | 22 | 0 | 308.114 | 0 | -211.99961 | hstg__passthrough_32 | 0 | 333.11361 | 0 | 358.114 | probe__eeg_mouse_1x30 | 0 | 20 | uV | 0 | 0 | 0 | 0 | 1 | -187 | -237 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_20 | E21 | pri_4 | ai0 | 20 | port_A_20 | 20 | circle | A | 23 | 0 | 308.114 | 0 | -349.99961 | hstg__passthrough_32 | 0 | 333.11361 | 0 | 358.114 | probe__eeg_mouse_1x30 | 0 | 21 | uV | 0 | 0 | 0 | 0 | 1 | -325 | -375 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_21 | E22 | pri_2 | ai0 | 21 | port_A_21 | 21 | circle | A | 19 | 0 | 456.114 | 0 | -111.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 22 | uV | 0 | 0 | 0 | 0 | 1 | -87 | -137 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 5 |
| pri_22 | E23 | pri_0 | ai0 | 22 | port_A_22 | 22 | circle | A | 24 | 0 | 456.114 | 0 | -287.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 23 | uV | 0 | 0 | 0 | 0 | 1 | -263 | -313 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_23 | E24 | pri_15 | ai0 | 23 | port_A_23 | 23 | circle | A | 25 | 0 | 456.114 | 0 | -404.99961 | hstg__passthrough_32 | 0 | 481.11361 | 0 | 506.114 | probe__eeg_mouse_1x30 | 0 | 24 | uV | 0 | 0 | 0 | 0 | 1 | -380 | -430 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_24 | E25 | pri_13 | ai0 | 24 | port_A_24 | 24 | circle | A | 20 | 0 | 564.114 | 0 | -112.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 25 | uV | 0 | 0 | 0 | 0 | 1 | -88 | -138 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 5 |
| pri_25 | E26 | pri_11 | ai0 | 25 | port_A_25 | 25 | circle | A | 26 | 0 | 564.114 | 0 | -287.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 26 | uV | 0 | 0 | 0 | 0 | 1 | -263 | -313 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_26 | E27 | pri_9 | ai0 | 26 | port_A_26 | 26 | circle | A | 27 | 0 | 564.114 | 0 | -412.99961 | hstg__passthrough_32 | 0 | 589.11361 | 0 | 614.114 | probe__eeg_mouse_1x30 | 0 | 27 | uV | 0 | 0 | 0 | 0 | 1 | -388 | -438 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_27 | E28 | pri_7 | ai0 | 27 | port_A_27 | 27 | circle | A | 21 | 0 | 674.114 | 0 | -99.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 28 | uV | 0 | 0 | 0 | 0 | 1 | -75 | -125 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 5 |
| pri_28 | E29 | pri_5 | ai0 | 28 | port_A_28 | 28 | circle | A | 28 | 0 | 674.114 | 0 | -223.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 29 | uV | 0 | 0 | 0 | 0 | 1 | -199 | -249 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
| pri_29 | E30 | pri_3 | ai0 | 29 | port_A_29 | 29 | circle | A | 29 | 0 | 674.114 | 0 | -404.99961 | hstg__passthrough_32 | 0 | 699.11361 | 0 | 724.114 | probe__eeg_mouse_1x30 | 0 | 30 | uV | 0 | 0 | 0 | 0 | 1 | -380 | -430 | 0 | 0 | SmartboxPro Signals | hstg__passthrough_32/probe__eeg_mouse_1x30 | 0 | 6 |
write_csv(stage2, "MouseEEGv2H32_Import_Stage2.csv")This table will be used to assign metadata to our final channel mapping and can be downloaded here.
This import process has been now incorporated into our vHTP import utility.
We’ll also post any updates from Neuronexus regarding this process and finding the reference channel.